GI-MAP | GI Microbial Assay Plus
Optimal Health — It All Starts with the GI-MAP™
New Add-Ons to GI-MAP Enhance Insight into Patient Health
August 2023 Update!
Fecal Gluten Peptide (Stool Gluten)
Fecal Gluten Peptide quantitatively evaluates the gluten immunogenic peptide in stool after recent ingestion (within two-four days). Results can guide practitioners to the source of "hidden" gluten exposures – allowing for increased adherence to a proper therapeutic diet.
The Gluten Peptide (Stool Gluten) marker is available as an optional add-on to the GI-MAP, or as a stand-alone test.
FECAL GLUTEN PEPTIDE - MORE INFOUniversal Antibiotic Resistance (AR) Genes Panel
The Universal Antibiotic Resistance Genes Panel detects the presence of 55 genetic elements associated with resistance to 10 different classes of antibiotics. Useful for patients who have been hospitalized, treated with antibiotics, or who have stubborn, chronic infections, the Universal AR Genes panel can inform practitioners if antibiotic resistance is a potential stumbling block for patients.
The Universal Antibiotic Resistance (AR) Gene Panel is available as an add-on only to the GI-MAP, or Pathogens Panel.
AR GENES - MORE INFOGI-MAP (Microbial Assay Plus) – View Sample Report
Please Note: Add-on options are available to order in the Results Portal.
View Sample ReportThe Industry's Leading Comprehensive Stool Test Just Got Better!
The Industry's Leading Comprehensive Stool Test Just Got Better!
The Industry's Leading Comprehensive Stool Test Just Got Better!
Features of the Updated GI-MAP Report
- New color reporting for easier visual readability
- Recategorized dysbiosis microbes for clear and concise test interpretation
- New markers added, including Roseburia spp. Desulfovibrio spp. and Eosinophil Activation Protein (EPX/EDN)
GI-MAP (Microbial Assay Plus) – View Sample Report
Please Note: Changes go into effect on samples received by the laboratory August 22, 2022 or later. Samples received before this date will not contain these updates.
View Sample ReportIMPORTANT: SARS-CoV-2 Stool Analysis Is Not Included on the GI-MAP and Must Be Ordered by Phone at (877) 485-5336. DO NOT use the GI-MAP kit for SARS-CoV-2 testing, it IS NOT the same test. See: SARS-CoV-2 Stool Analysis for additional information.
Results You Can Rely On
Why Is Quantification Using qPCR Technology So Important?
Unlike other comprehensive stool tests on the market, the GI-MAP can provide practitioners with truly quantitative results. qPCR offers a much more accurate way to detect and quantify clinically-relevant organisms than standard PCR, culture, microscopy, or DNA sequencing-based methods. Accurately assessing how much of an organism's DNA is present in a patient's stool sample is essential for helping practitioners to determine the clinical significance of pathogenic organisms and dysbiosis patterns.
qPCR's Reliability, Reproducibility, and Use in Clinical Research
 Although qPCR is becoming more commonplace in in-vitro diagnostics (IVD), we are the only laboratory in the United States exclusively using qPCR technology for advanced comprehensive stool testing. This technology is used routinely in clinical and academic research because it provides highly-accurate quantification, as well as high levels of sensitivity and specificity. Standard PCR technology doesn't offer the same level of sensitivity, or the ability to express precise numerical results.
Although qPCR is becoming more commonplace in in-vitro diagnostics (IVD), we are the only laboratory in the United States exclusively using qPCR technology for advanced comprehensive stool testing. This technology is used routinely in clinical and academic research because it provides highly-accurate quantification, as well as high levels of sensitivity and specificity. Standard PCR technology doesn't offer the same level of sensitivity, or the ability to express precise numerical results.
The GI-MAP also provides consistently reproducible results. Reproducibility is of crucial importance to the practitioners and patients that rely on the efficacy of the GI-MAP. To achieve it, we perform rigorous quality control, and have validated all molecular target quantification assays to meet or exceed FDA standards.
GI-MAP Allows for the Personalized Treatment Plans and Informative Retests
The GI-MAP's accuracy and reliability allows practitioners to create personalized treatment protocols to address gut dysfunction based on which infections are urgent, which areas of the gut are already optimized, and which areas should be addressed after an infection is resolved.
Additionally, the quantification offers a remarkable ability to see how treatment modalities are working because a retest after treatment can show whether a parasite has resolved, dysbiosis has improved, and more.
Who Should Have the GI-MAP Comprehensive Stool Analysis Done?

Almost every patient can benefit from a GI-MAP gut health assessment. Some patients are looking to achieve optimal health, while other patients have been chronically ill and frustrated without a diagnosis for years.
Some conditions that warrant testing are:
- Autoimmune diseases
- IBS/IBD
- Digestive complaints, diarrhea or constipation
- Brain fog
- Skin problems, like acne and psoriasis
- Mood disorders, depression, and anxiety
- Diabetes and weight loss issues
Can Infants and Children Benefit from the GI-MAP?
Yes. The GI-MAP is commonly used in infants and children, and can provide insight into conditions related to Attentional Deficit Hyperactivity Disorder, Autism, and digestive complaints. Patients – See Ordering Info Below
What is Reported on GI-MAP Test Results?
The Gastrointestinal Microbial Assay Plus (GI-MAP) is an innovative clinical tool that measures gastrointestinal microbiota DNA from a single stool sample with state of the art, quantitative polymerase chain reaction (qPCR) technology.
The GI-MAP was designed to detect microbes that may be disturbing normal microbial balance or contributing to illness as well as indicators of digestion, absorption, inflammation, and immune function.
Page by Page Listing of Microorganisms Found on the GI-MAP:
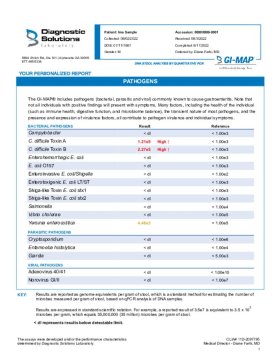
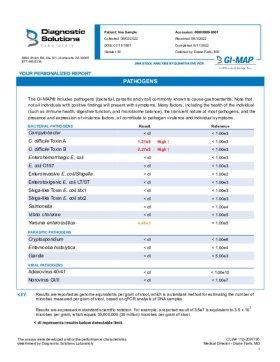
PATHOGENS
The GI-MAP includes pathogens (bacterial, parasitic, and viral) commonly known to cause intestinal gastroenteritis. It's important to note that not all individuals with positive findings for pathogens will present with symptoms. Many factors, including the health of the individual, the transient nature of some pathogens, and the presence and expression of virulence factors all contribute to an individual's symptoms.
Toxins are a type of virulence factor produced by certain pathogens. Since GI-MAP is a DNA-based test, results reflect the levels of pathogenic strains carrying the toxin genes, not the levels of any toxins that may be produced.
BACTERIAL PATHOGENS
- Campylobacter
- C. difficile Toxin A
- C. difficile Toxin B
- Enterohemorrhagic E. coli
- E. coli O157
- Enteroinvasive E. coli/Shigella
- Enterotoxigenic E. coli LT/ST
- Shiga-like Toxin E. coli stx1
- Shiga-like Toxin E. coli stx2
- Salmonella
- Vibro cholerae
- Yersinia enterocolitica
PARASITIC PATHOGENS
- Cryptosporidium
- Entamoeba histolytica
- Giardia
VIRAL PATHOGENS
- Adrenovirus 40/41
- Norovirus GI/II
H. pylori
Recent studies have shown that nearly 50% of the world's population may harbor H. pylori. And, although many carriers are asymptomatic, H. pylori is known to have a causative role in ulcers, chronic gastritis, and stomach cancer.
Additionally, in early phases of colonization, patients may experience hypochlorhydria followed by a change to hyper aciduria. Over time, additional H. pylori strains may colonize, including those with Virulence Factors and increased disease potential.
H. PYLORI & VIRULENCE FACTORS
- Helicobacter pylori
- Virulence Factor, babA
- Virulence Factor, cagA
- Virulence Factor, dupA
- Virulence Factor, iceA
- Virulence Factor, oipA
- Virulence Factor, vacA
- Virulence Factor, virB
- Virulence Factor, virD
COMMENSAL/KEYSTONE BACTERIA
Trillions of microorganisms inhabit the human intestine to make up a complex ecosystem that plays an important role in human health. Commensal bacteria extract nutrients and energy from our diets, maintain gut barrier function, produce vitamins (biotin and vitamin K), and protect against colonization by potential pathogens.
COMMENSAL BACTERIA
- Bacteroides fragilis
- Bifidobacterium spp.
- Enterococcus spp.
- Escherichia spp.
- Lactobacillus spp.
- Enterobacter spp.
- Akkermansia muciniphilia
- Faecalbacterium prausnitzii
- Roseburia spp.
BACTERIAL PHYLA
- Bacteroidetes
- Firmicutes
- Firmicutes/Bacteroidetes Ratio
OPPORTUNISTIC/OVERGROWTH MICROBES
Many bacteria measured on the GI-MAP are considered opportunistic pathogens, as they only cause disease and illness in some individuals, particularly the immune-compromised. Many individuals come into contact with opportunistic bacteria and experience no symptoms. Most sources consider these microbes to be normal in the stool. However, they can cause gastroenteritis and inflammation at high levels in vulnerable patients. Symptoms may include diarrhea, loose stools, abdominal pain, or even constipation.
Overgrowth and excessive colonization by opportunistic bacteria may occur when the commensal bacteria are impaired by poor diet, antibiotic use, parasitic infection, or a weakened immune system. When intestinal permeability is present (see zonulin), these microbes could escape the lumen of the gut and infect extraintestinal sites.
OPPORTUNISTIC/OVERGROWTH MICROBES
- Bicillus spp.
- Enterococcus faecalis
- Enterococcus faecium
- Morganella spp.
- Pseudomonas spp.
- Pseudomonas aeruginosa
- Staphylococcus spp.
- Staphylococcus aureus
- Streptococcus spp.
COMMENSAL OVERGROWTH MICROBES
- Desulfovibrio spp.
- Methanobacteriaceae (family)
INFLAMMATORY & AUTOIMMUNE-RELATED BACTERIA
- Citrobacter spp.
- Citrobacter freundii
- Klebsiella spp.
- Klebsiella pneumoniae
- M. avium subsp. paratuberculosis
- Proteus spp.
- Proteus mirabilis
COMMENSAL INFLAMMATORY & AUTOIMMINE-RELATED BACTERIA
- Enterobacter spp.
- Escherichia spp.
- Fusobacterium spp.
- Prevotella copri
FUNGI/YEAST
Fungal organisms are commonly found in the human digestive tract, but fungal overgrowth can cause illness in susceptible individuals. Fungal growth may be localized in the body. For instance, Candida spp. may be high in the large intestine but normal in the small intestine, and vice versa. In a patient with suspected fungal overgrowth, additional tests may be necessary to understand the complete picture of fungal overgrowth. Urinary D-arabinitol or antibodies to Candida are sometimes used.
FUNGI/YEAST
- Candida spp.
- Candida albicans
- Geotricum spp.
- Microsporidia spp.
- Rhodoturula spp.
VIRUSES
Copy?
VIRUSES
- Cytomegalovirus
- Epstein Bar Virus
PARASITES
A parasite is an organism that lives and feeds on a host organism at the expense of the host. The GI-MAP tests for pathogenic parasites and protozoa (some of which are non-pathogenic) most commonly occurring in the GI tract. Sources of exposure should be identified and eliminated to prevent reinfection.
PROTOZOA
- Blastocystis hominis
- Chilomastix mesnelli
- Cyclospora spp.
- Dientamoeba fragilis
- Endolimax nana
- Entamoeba coli
- Pentatrichomonas hominis
WORMS
- Ancyclostroma duodenale
- Ascaris lumbricoides
- Necator americanis
- Trichuris trichiura
- Taenia spp.
INTESTINAL HEALTH MARKERS
Copy?
DIGESTION
- Steatocrit
- Elastase-1
GI MARKERS
- β-Glucuronidase
- Occult Blood – FIT
IMMUNE RESPONSE
- Secretory IgA
- Anti-gliadin IgA
- Eosinophil Activation Protein (EDN)
INFLAMMATION
- Calprotectin
ADD-ON TESTS
- Zonulin
- Gluten Peptide
- Universal Antibiotic Resistance Genes – This option is reported on page 6.
H. PYLORI ANTIBIOTIC RESISTANCE GENES
The GI-MAP includes results for detection of antibiotic resistance genes in the microbiome. If an antibiotic resistance gene is present, then that class of antibiotics is designated POSITIVE for antibiotic resistance. A positive result for the presence of resistance genes for a given antibiotic indicates that the antibiotic is not an ideal choice for an antibiotic protocol.
Antibiotic resistance genes apply to all of the microorganisms found in the fecal sample. Since microbes can rapidly share DNA under stress, the presence of antibiotic resistance in any organism is reason enough to avoid that drug class.
Phenotypes | HELOBACTER
- Amoxicillen
- Clarithromycin
- Fluroquinolines
- Tetracycline
Genotypes | UNIVERSAL MICROBIOTA RESISTANCE GENES
- β-lactamase
- Fluoroquinolones
- Macrolides
- Vancomycin
UNIVERSAL MICROBIOTA RESISTANCE GENES
- β-lactamase
- Fluoroquinolones
- Macrolides
- Vancomycin
UNIVERSAL ANTIBIOTIC RESISTANCE GENES
The Universal Antibiotic Resistance (AR) Genes panel can be ordered as an add-on to the GI-MAP or to the GI-Pathogens panel. It detects the presence of 55 genetic elements associated with resistance to 10 different classes of antibiotics.
This AR Genes add-on panel detects genes associated with resistance to the most commonly prescribed antibiotics for gastrointestinal infections. The mobile genetic elements in the panel can be found in a variety of different microbes. The presence of these genes in a bacterial population have been associated with moderate to high levels of antibiotic resistance in human gastrointestinal infections.
UNIVERSAL ANTIBIOTIC RESISTANCE GENES
- β-lactamase
- Fluoroquinolones
- Vancomycin
- Macrolides
- Ciprofloxacin
- 5-Nitroimidazoles (non-Helicobacter pylori)
- Trimethoprim
- Sulfonamides
- Methicillin
- Chloramphenicol
Genotypes | UNIVERSAL MICROBIOTA RESISTANCE GENES
- β-lactamase
- Fluoroquinolones
- Macrolides
- Vancomycin
UNIVERSAL MICROBIOTA RESISTANCE GENES
- β-lactamase
- Fluoroquinolones
- Macrolides
- Vancomycin
What is Reported on GI-MAP Test Results?
The Gastrointestinal Microbial Assay Plus (GI-MAP) is an innovative clinical tool that measures gastrointestinal microbiota DNA from a single stool sample with state of the art, quantitative polymerase chain reaction (qPCR) technology.
The GI-MAP was designed to detect microbes that may be disturbing normal microbial balance or contributing to illness as well as indicators of digestion, absorption, inflammation, and immune function.
Page by Page Listing of Microorganisms Found on the GI-MAP:
PATHOGENS
The GI-MAP™ includes pathogens (bacterial, parasitic, and viral) commonly known to cause intestinal gastroenteritis. It's important to note that not all individuals with positive findings for pathogens will present with symptoms. Many factors, including the health of the individual, the transient nature of some pathogens, and the presence and expression of virulence factors all contribute to an individual's symptoms.
Toxins are a type of virulence factor produced by certain pathogens. Since GI-MAP is a DNA-based test, results reflect the levels of pathogenic strains carrying the toxin genes, not the levels of any toxins that may be produced.
BACTERIAL PATHOGENS
- Campylobacter
- C. difficile Toxin A
- C. difficile Toxin B
- Enterohemorrhagic E. coli
- E. coli O157
- Enteroinvasive E. coli/Shigella
- Enterotoxigenic E. coli LT/ST
- Shiga-like Toxin E. coli stx1
- Shiga-like Toxin E. coli stx2
- Salmonella
- Vibro cholerae
- Yersinia enterocolitica
PARASITIC PATHOGENS
- Cryptosporidium
- Entamoeba histolytica
- Giardia
VIRAL PATHOGENS
- Adrenovirus 40/41
- Norovirus GI/II
H. pylori
Recent studies have shown that nearly 50% of the world's population may harbor H. pylori. And, although many carriers are asymptomatic, H. pylori is known to have a causative role in ulcers, chronic gastritis, and stomach cancer.
Additionally, in early phases of colonization, patients may experience hypochlorhydria followed by a change to hyper aciduria. Over time, additional H. pylori strains may colonize, including those with Virulence Factors and increased disease potential.
H. PYLORI & VIRULENCE FACTORS
- Helicobacter pylori
- Virulence Factor, babA
- Virulence Factor, cagA
- Virulence Factor, dupA
- Virulence Factor, iceA
- Virulence Factor, oipA
- Virulence Factor, vacA
- Virulence Factor, virB
- Virulence Factor, virD
COMMENSAL/KEYSTONE BACTERIA
Trillions of microorganisms inhabit the human intestine to make up a complex ecosystem that plays an important role in human health. Commensal bacteria extract nutrients and energy from our diets, maintain gut barrier function, produce vitamins (biotin and vitamin K), and protect against colonization by potential pathogens.
COMMENSAL BACTERIA
- Bacteroides fragilis
- Bifidobacterium spp.
- Enterococcus spp.
- Escherichia spp.
- Lactobacillus spp.
- Enterobacter spp.
- Akkermansia muciniphilia
- Faecalbacterium prausnitzii
- Roseburia spp.
BACTERIAL PHYLA
- Bacteroidetes
- Firmicutes
- Firmicutes/Bacteroidetes Ratio
OPPORTUNISTIC/OVERGROWTH MICROBES
Many bacteria measured on the GI-MAP are considered opportunistic pathogens, as they only cause disease and illness in some individuals, particularly the immune-compromised. Many individuals come into contact with opportunistic bacteria and experience no symptoms. Most sources consider these microbes to be normal in the stool. However, they can cause gastroenteritis and inflammation at high levels in vulnerable patients. Symptoms may include diarrhea, loose stools, abdominal pain, or even constipation.
Overgrowth and excessive colonization by opportunistic bacteria may occur when the commensal bacteria are impaired by poor diet, antibiotic use, parasitic infection, or a weakened immune system. When intestinal permeability is present (see zonulin), these microbes could escape the lumen of the gut and infect extraintestinal sites.
OPPORTUNISTIC/OVERGROWTH MICROBES
- Bicillus spp.
- Enterococcus faecalis
- Enterococcus faecium
- Morganella spp.
- Pseudomonas spp.
- Pseudomonas aeruginosa
- Staphylococcus spp.
- Staphylococcus aureus
- Streptococcus spp.
COMMENSAL OVERGROWTH MICROBES
- Desulfovibrio spp.
- Methanobacteriaceae (family)
INFLAMMATORY & AUTOIMMUNE-RELATED BACTERIA
- Citrobacter spp.
- Citrobacter freundii
- Klebsiella spp.
- Klebsiella pneumoniae
- M. avium subsp. paratuberculosis
- Proteus spp.
- Proteus mirabilis
COMMENSAL INFLAMMATORY & AUTOIMMINE-RELATED BACTERIA
- Enterobacter spp.
- Escherichia spp.
- Fusobacterium spp.
- Prevotella copri
FUNGI/YEAST
Fungal organisms are commonly found in the human digestive tract, but fungal overgrowth can cause illness in susceptible individuals. Fungal growth may be localized in the body. For instance, Candida spp. may be high in the large intestine but normal in the small intestine, and vice versa. In a patient with suspected fungal overgrowth, additional tests may be necessary to understand the complete picture of fungal overgrowth. Urinary D-arabinitol or antibodies to Candida are sometimes used.
FUNGI/YEAST
- Candida spp.
- Candida albicans
- Geotricum spp.
- Microsporidia spp.
- Rhodoturula spp.
VIRUSES
Copy?
VIRUSES
- Cytomegalovirus
- Epstein Bar Virus
PARASITES
A parasite is an organism that lives and feeds on a host organism at the expense of the host. The GI-MAP tests for pathogenic parasites and protozoa (some of which are non-pathogenic) most commonly occurring in the GI tract. Sources of exposure should be identified and eliminated to prevent reinfection.
PROTOZOA
- Blastocystis hominis
- Chilomastix mesnelli
- Cyclospora spp.
- Dientamoeba fragilis
- Endolimax nana
- Entamoeba coli
- Pentatrichomonas hominis
WORMS
- Ancyclostroma duodenale
- Ascaris lumbricoides
- Necator americanis
- Trichuris trichiura
- Taenia spp.
INTESTINAL HEALTH MARKERS
Copy?
DIGESTION
- Steatocrit
- Elastase-1
GI MARKERS
- β-Glucuronidase
- Occult Blood – FIT
IMMUNE RESPONSE
- Secretory IgA
- Anti-gliadin IgA
- Eosinophil Activation Protein (EDN)
INFLAMMATION
- Calprotectin
ADD-ON TESTS
- Zonulin
H. PYLORI ANTIBIOTIC RESISTANCE GENES
The GI-MAP includes results for detection of antibiotic resistance genes in the microbiome. If an antibiotic resistance gene is present, then that class of antibiotics is designated POSITIVE for antibiotic resistance. A positive result for the presence of resistance genes for a given antibiotic indicates that the antibiotic is not an ideal choice for an antibiotic protocol.
Antibiotic resistance genes apply to all of the microorganisms found in the fecal sample. Since microbes can rapidly share DNA under stress, the presence of antibiotic resistance in any organism is reason enough to avoid that drug class.
Phenotypes | HELOBACTER
- Amoxicillen
- Clarithromycin
- Fluroquinolines
- Tetracycline
Genotypes | UNIVERSAL MICROBIOTA RESISTANCE GENES
- β-lactamase
- Fluoroquinolones
- Macrolides
- Vancomycin
UNIVERSAL MICROBIOTA RESISTANCE GENES
- β-lactamase
- Fluoroquinolones
- Macrolides
- Vancomycin
Start Your GI-MAP Journey Today
Teaming up with us is easy using our Account Setup page. Testing is backed up by scientific literature and uses advanced testing methods that provide reliable quantitative results with clinically-actionable decision points for your patients. (See information below if you're a patient.)
The GI-MAP is the most accurate, comprehensive DNA stool analysis on the market. Become part of our team of practitioners who regularly use it to improve patient outcomes. (See what practitioners are saying about the GI-MAP stool test.)
Our commitment to laboratory medicine is to utilize proven methodologies that are accurate and reliable.
Patients

Are You a Patient?
A healthcare practitioner will need to order the test for you. That can be your own doctor, or we can help you find a practitioner in your area.
We know that other companies may offer to sell stool testing directly to you. The GI-MAP is a diagnostic tool and relies on state-of-the-art technology to provide insights into your health. It can be used for patients suffering from serious illnesses, like autoimmune diseases, thyroid diseases, parasitic infections, and Crohn's Disease – to name a few.
These serious health concerns should be treated by a licensed practitioner who can interpret your test results and tailor a treatment plan unique to your specific needs.
Your health is your greatest asset – only entrust it to an expert.
Our transparent patient billing system is easy to use, and we file claims to insurance and Medicare.
Why We're So Passionate about Patient Health
Since the introduction of the GI-MAP, we have received hundreds of messages from practitioners and patients about how a simple stool test (that can be collected in one day), has changed their lives for the better. Ultimately, improved patient outcomes are our goal.
Every test that comes through our door is an opportunity for us to deliver life-changing results to practitioners that are working with patients that need answers. Sometimes those results show that a patient is suffering from multiple parasites. When a gut infection is found it presents an opportunity for the healing journey to begin and for patients to win. And, that's why we do what we do.
Diagnostic Solutions Laboratory is CLIA Certified
In addition to our internal quality control and validation processes, Diagnostic Solutions Laboratory is CLIA certified. CLIA certification ensures that our laboratory meets or exceeds U.S. standards set for laboratory excellence.
- Support item one
- Support item one
- Support item one
The GI-MAP is Constantly Evolving

The Human Microbiome Project began in 2007. Since then, the amount of research about the inhabitants of the gut microbiome and the impact they have on health has expanded exponentially. When new bacterial, commensal, or other organisms of relevance are discovered, we begin working on adding them to the GI-MAP. Our test moves in the same direction as the science that comes out about it does.
Are You a New GI-MAP User?
Get all the basics by watching this informative GI-MAP Overview
FEATURED VIDEO
GI-MAP is Revolutionizing the Comprehensive Stool Analysis
Microscopy and culture-based tests are familiar, but they have limitations with sensitivity, specificity, and identifying anaerobic organisms – that is why almost no peer-reviewed research studies have used microscopy and culture-based testing methods in over 25 years.
Methodology
- Quantitative PCR (qPCR) or Real-Time Polymerase Chain Reaction (RT-PCR) – Provides you with true quantitative values. It helps differentiate trace levels of an organism from frank elevations indicative of active infection.
Specimen Requirements
- Single Stool Sample – Ambient room temperature in specimen vial provided.

Test Ordering Options
There are two testing options for :
- GI-MAP™ – GI Microbial Assay Plus
- GI-MAP™ + Zonunlin
- GI Pathogens Profile
- H. pylori Profile
- Zonulin Profile (Stool)
- Calprotectin
- Gluten Peptide (Stool Gluten)
- Universal Antibiotic Resistance Genes Panel
If you already have an account with us, you may order through our online Results Portal accessible at the top of this page.

Complementary Panels for Complex Cases
- OMX Organic Metabolomics – Comprehensive metabolic profiling that evaluates patterns of metabolites related to core biological systems, offering insight into biochemical dysfunctions that may be of concern.
- GenomicInsight – Data gleaned from GenomicInsight informatics uses the latest medical literature to provide relevant information on nutraceuticals, nutritional supplements, diet, and lifestyle interventions that can proactively influence a patient's SNPs to reduce or prevent disease risk. Furthermore, pharmacogenomic results included in the profile allow practitioners to predict the efficacy of select pharmaceuticals tailored the individual's genetic make-up.
Information provided by Diagnostic Solutions Laboratory does not constitute medical advice; but is for educational purposes only.
Services provided are for laboratory testing only. No charge is incurred for ordering collection kits.
8.17.2023
Sample Report
Product Brief
GI-MAP Interpretive Guide
This guide has been updated to include changes to the report introduced August, 2023.
GI-MAP Articles
Fecal Gluten Peptide – Published August 10, 2023
Antibiotic Resistance Genes – Published August 11, 2023
Gas & Histamine Producers on GI-MAP – Published January 31, 2022
qPCR Testing for GI Pathogens – Published January 13, 2021
Response to DDI Funded Study – Published September 30, 2020
Strengthening the Immune System via GI Infection Treatments – Published April 16, 2020
Botanical Therapy for Antibiotic Resistance in the GI Tract – Published November 20, 2019
Autoimmune Diseases: How to Get to the Bottom of the Problem – Published September 17, 2019
GI Testing is Imperative, Even in Patients Without Digestive Complaints – Published May 15, 2019
The GI-MAP in Clinical Practice – Published September 6, 2018
Technology Update: Advances to the GI-MAP – Published March 13, 2018
GI-MAP — Now Even Better! – Published December 20, 2017
Newsletter Sign-up
By submitting this form, you are consenting to receive marketing emails from: Diagnostic Solutions Laboratory. You can revoke your consent to receive emails at any time by using the SafeUnsubscribe® link, found at the bottom of every email.